Welcome to RegRegSEA!
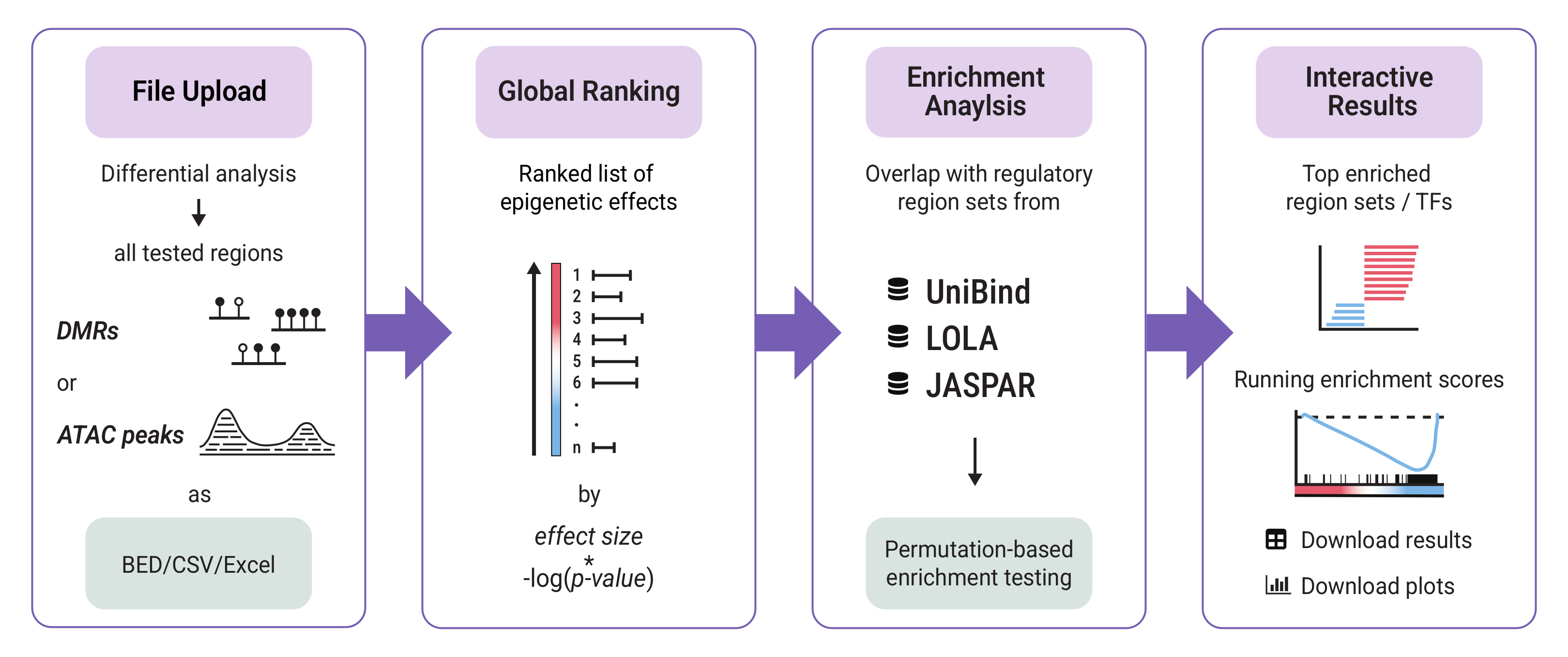
RegRegSEA is a webserver designed for Regulatory Region Set Enrichment Analysis of epigenomic data. By adapting the statistical approach of GSEA to the epigenetic landscape, our tool allows you to test for the enrichment of regulatory region sets in DNA methylation and chromatin accessibility data.
Instead of analysing a truncated list of statistically significant effects requiring strict cutoffs, RegRegSEA evaluates the entire ranked list of genomic intervals. This sensitive approach allows for identification of transcription factors and regulatory elements associated with subtle but coordinated global shifts in chromatin accessibility or DNA methylation across the genome.
Supported regulatory region set databases:
- UniBind TFBSs – Experimentally validated TFBSs from ChIP-seq data
- LOLA Region Databases – TFBSs, histone modifications and broader genomic features
- JASPAR – TF motif matches
This webserver is free and open to all users and does not require a login.
Quick Actions
Citation
If you use RegRegSEA in your research, please cite:
RegRegSEA: A Webserver for Regulatory Region Set Enrichment Analysis of Epigenomic Data
as well as the respective region set databases used in your analysis.
Getting Started
- Prepare your data: Export differential analysis results with genomic coordinates (chr, start, end), effect sizes, and raw p-values for all tested regions.
- Select databases: Choose regulatory region set databases (UniBind, LOLA, JASPAR) relevant to your research.
- Submit and monitor: Upload data via New Submission and track progress via a job ID on the Results page.
- Explore results: View enrichment tables, visualize top region sets and running enrichment scores, and download results and leading edge regions (BED) for downstream analyses (e.g. integration with gene expression data).
Use Cases
DNA Methylation Studies
Identify transcription factors associated with DNA methylation changes in disease, development, or environmental response.
Chromatin Accessibility Studies
Discover regulatory elements associated with alterations in chromatin accessibility.
Multi-Omics Integration
Connect epigenetic alterations to transcriptional regulators and gene expression programs.